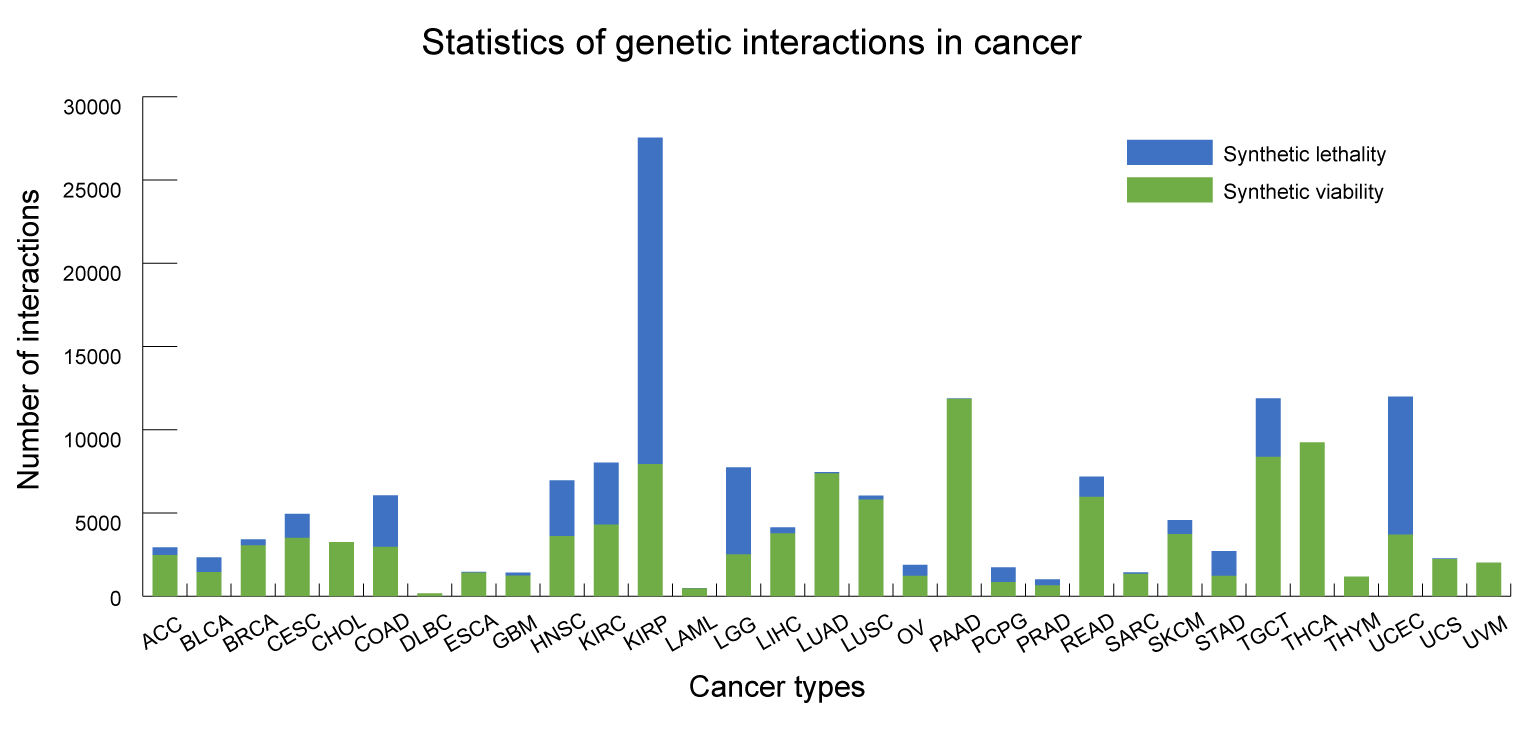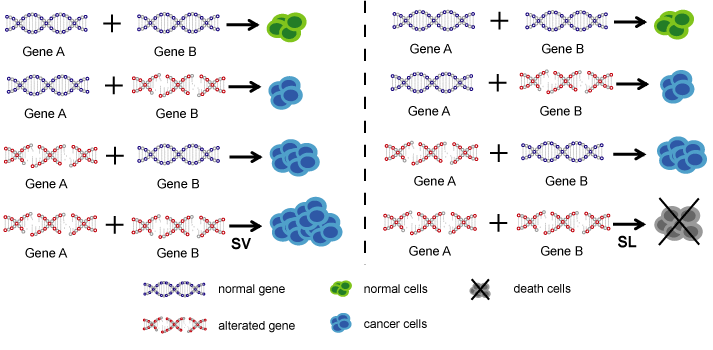
Cancer cells generally harbor hundreds of alterations in the cancer genomes and act as crucial factors in the development and progression of cancer. Gene alterations in the cancer genome form genetic interactions, which affect the response of patients to drugs. The generation of an unexpected phenotypic outcome when combining two alterations is referred to as a genetic interaction, including synthetic viability and synthetic lethality. Synthetic viability (SV) describes the scenario in which the synthesis or combination of two gene effects remedy the effect of single-gene defects, such as cell death or a significant impairment of fitness, and enhance cell survival, which may be a potential mechanism for drug resistance. Synthetic lethality (SL) describes the scenario in which single-gene defects are compatible with cell viability, but the combination of alterations in both genes induces cell death, which is generally used to identify drug sensitive biomarkers.
However, detection of genetic interactions in the cancer genome is a major obstacle in the research of identifying biomarkers for cancer cell drug resistance and sensitivity. In this database, an algorithm that mines copy number alteration and whole-exome mutation profiles from The Cancer Genome Atlas (TCGA), as well as functional screen data, such as CRISPR, was generated to identify potential genetic interactions for specific cancer types. By integrating current reported genetic interactions from other studies, the Cancer Genetic Interaction database (CGIdb) (http://www.medsysbio.org/CGIdb) was constructed, providing a convenient retrieval for genetic interactions in cancer. CGIdb provides the genetic interaction for specific cancer type, information of functional relationship between genetic interactions, the drug response (resistance or sensitivity) and so on. CGIdb provides a landscape of genetic interaction spectrum in cancer cells and reveals biomarkers related to cancer cell drug resistance and sensitivity, providing a reference for personalized treatment regimens.